MetaClass is a Karyotyping and FISH modular system used in human cytogenetic studies of chromosomes.
It consists in the following independent modules:
MetaClass Karyotyping MetaClass FISH MetaClass DatabaseMore information:
Metaclass Karyotyping
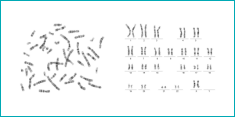
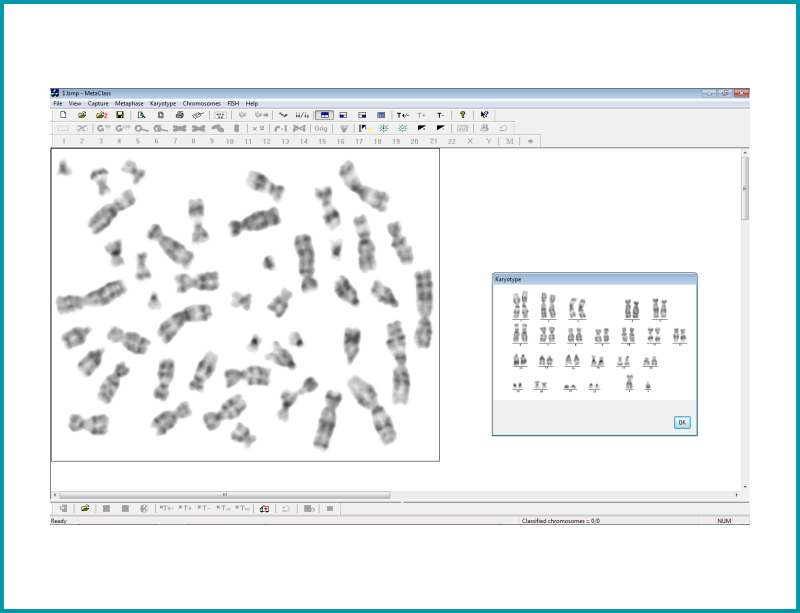
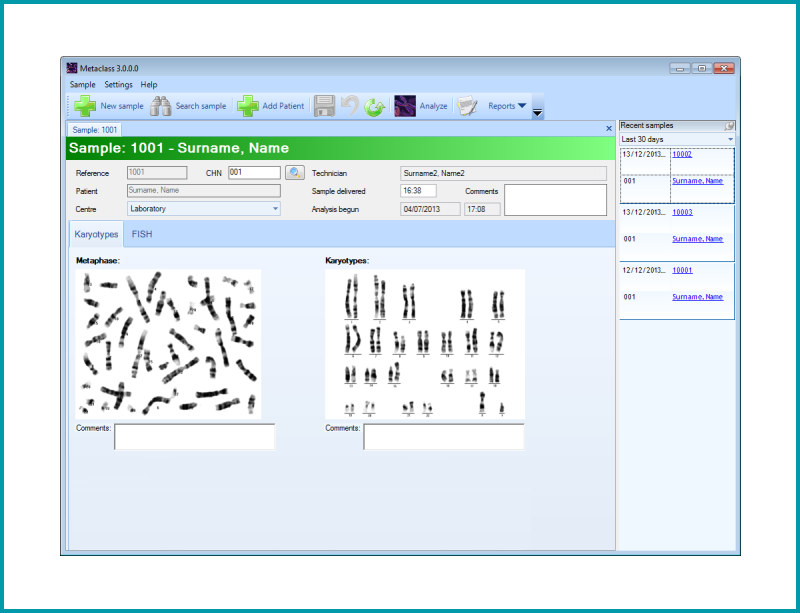
Allows the study of a karyotype following the capture of digital images from the metaphase preparation. The cytogeneticist can examine any structural change in the chromosomes. Each chromosome can be arranged in pairs according to size, position of the centromeres and banding pattern displayed.
It gives an excellent morphology and quality in the chromosome bands.
As final result the system will provides a personalized report.
| Karyotype study is analysed under brightfield microscopy. 100X magnification. |
| Several Metaphase images can be captured |
| Threshold can be modified to improve the captured image quality |
| Chromosomes are automatically classified |
| The module offers 3 views on the screen: Metaphase view, Karyotype and Unclassified chromosomes window marked with 0 |
| Chromosome prototype can be displayed |
| Unclassified chromosomes can be automatically corrected, or manually if desired, for the following: – Superimpositions: A chromosome appears superimposed to other in one part – Separation: When two chromosomes are joint in an extreme, can be automatically – Crossings – Straightening – Rotations – Flip a chromosome – Obtaining the specular image, as well as Magnification of the individual chromosomes is available – Best Brightness and contrast can be modified and automatically obtained. – Damaged chromosomes bands can be highlighted in the karyotype study. |
| *mtc session file or * bpm image can be saved and recalled. |
| Individual chromosomes can be selected and exported to the sample Management (Metaclass Database) to add comments to the final results and printable report. |
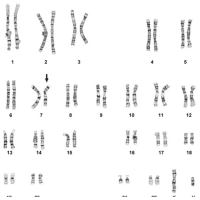
Down Syndrome karyotype (trisomy 21)
This is the most common numerical abnormality found in newborns. It is characterized by an extra chromosome 21 and the karyotype is written as: 47,XY,+21. The key to the karyotype description is as follows:
47: the total number of chromosomes (46 is normal). XY: the sex chromosomes (male). +21: designates the extra chromosome as a 21.Deletion 7 karyotype
This karyotype is an example of a simple deletion in one chromosome. In this case a segment within the q, or long arm of the right chromosome 7 is deleted. In this particular example there are two microscopically visible breaks within the long arm making it an interstitial deletion. If there had been one break resulting in the loss of the end of a chromosome this would be called a terminal deletion. The karyotype is written as: 46,XY,del(7)(q11.23q21.2). The key to the karyotype description is as follows:
46: the total number of chromosomes. XY: the sex chromosomes (male). del(7): deletion in chromosome 7. (q11.23q21.2): breakpoints of the deleted segment.Metaclass FISH
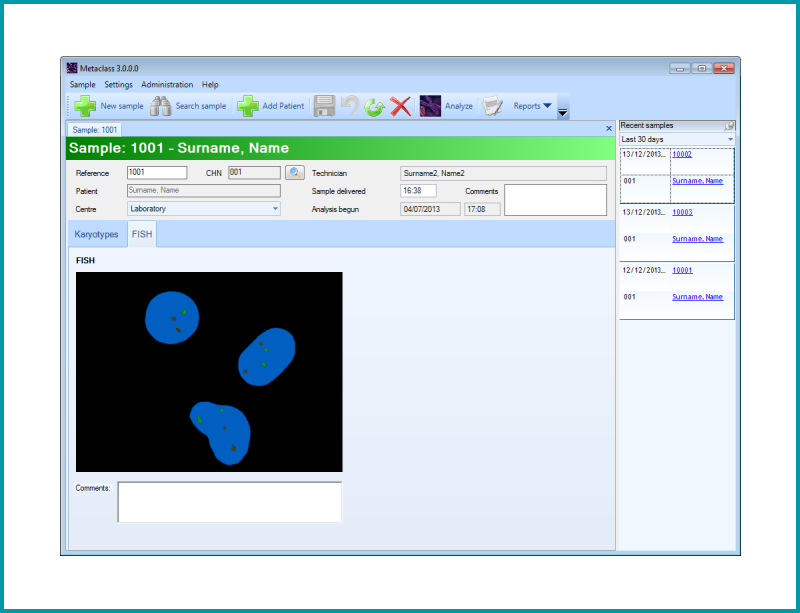
Analysis module to obtain the Fluorescent IN SITU Hybridization (FISH) image.
The images captured using fluorescently labelled DNA probes, allow the confirmation of genetic or chromosome abnormalities such as trisomies, microdelections or some chromosome rearrangements.
| FISH images are analysed under fluorescence microscopy, at 100X magnification |
| Several images are captured with different filters. The images are originally in b/w, but provide a unique color final image displaying the presence or absence of the signal. |
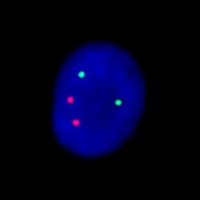
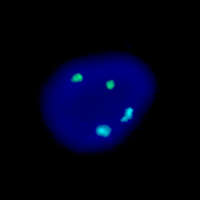
This is an example of the Aneuploid Screen test, where interphase nuclei from amniotic fluid cells have been combined with DNA probes for chromosomes 13, 18, 21, X, and Y. The nucleus on the left has been hybridized to probes for chromosomes 13 (green), and 21 (red). The nucleus on the right has been hybridized to probes for chromosomes 18 (aqua), X (green), and Y (red). Since there are two signals each for chromosomes 13, 18, and 21 and one signal each for the X and Y chromosome this fetus is a normal male with respect to the Aneuploid Screen test.
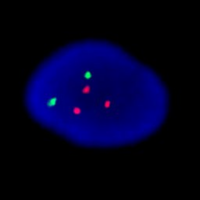
This Aneuploid Screen test shows a female fetus with trisomy-21. The nucleus on the left has been hybridized to probes for chromosomes 13 (green), and 21 (red) and clearly has three red signals. The nucleus on the right has been hybridized to probes for chromosomes 18 (aqua), X (green), and Y (red). Since it has two green signals and no red signal it is female.
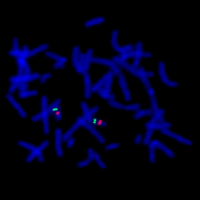
This is an example of a metaphase cell that has been hybridized with the probe for DiGeorge/Velo-Cardio-Facial/CATCH 22/Shprintzen Syndrome which is caused by a microdeletion on chromosome 22. The probe in this particular case is a dual-color mixture of two seperate probes for chromosome 22. The green signal is an internal control and is located at 22q13. It allows for quick identification of both #22 chromosomes. The red signal is located at the DiGeorge region at 22q11.2. Since both 22’s have the red signal in this cell there is not a microdeletion within the DiGeorge region and this individual would not have DiGeorge Syndrome.
MetaClass Database
Is an integrated database that can be installed with any of the above analysis modules, allowing the storage of patient details and giving an easy access to the results and final reports.
Customized reports in any language are included with this module.